scTimeBench
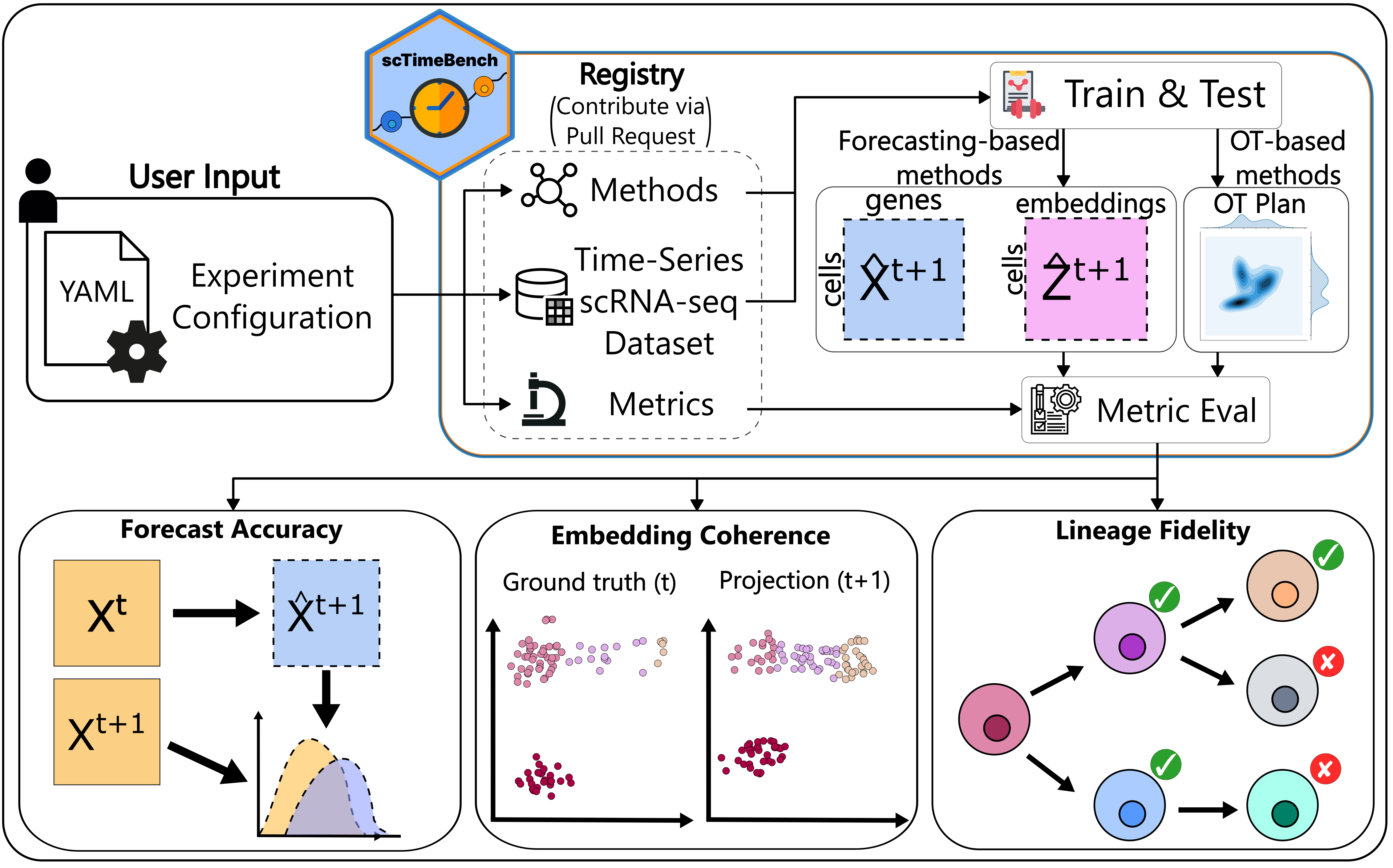
While building a method that is able to capture the trajectory of cells, we noticed that current single-cell time-series methods all benchmark solely on different distribution similarity metrics, but are:
- Not standardized across papers
- Only show cell lineage and cell-type information as an additional analysis, not something that needs to be verified and benchmarked.
This project resolves that by building something that I hope will be used by the community, and be maintained as new metrics, new analyses, new methods, and new datasets are being included. My main contribution to this project was ensuring that this framework is well-engineered, and easy to use! With my background in software engineering and working towards creating better user experience for machine learning researchers, I recognize what is needed to make sure using scTimeBench is a smooth experience.
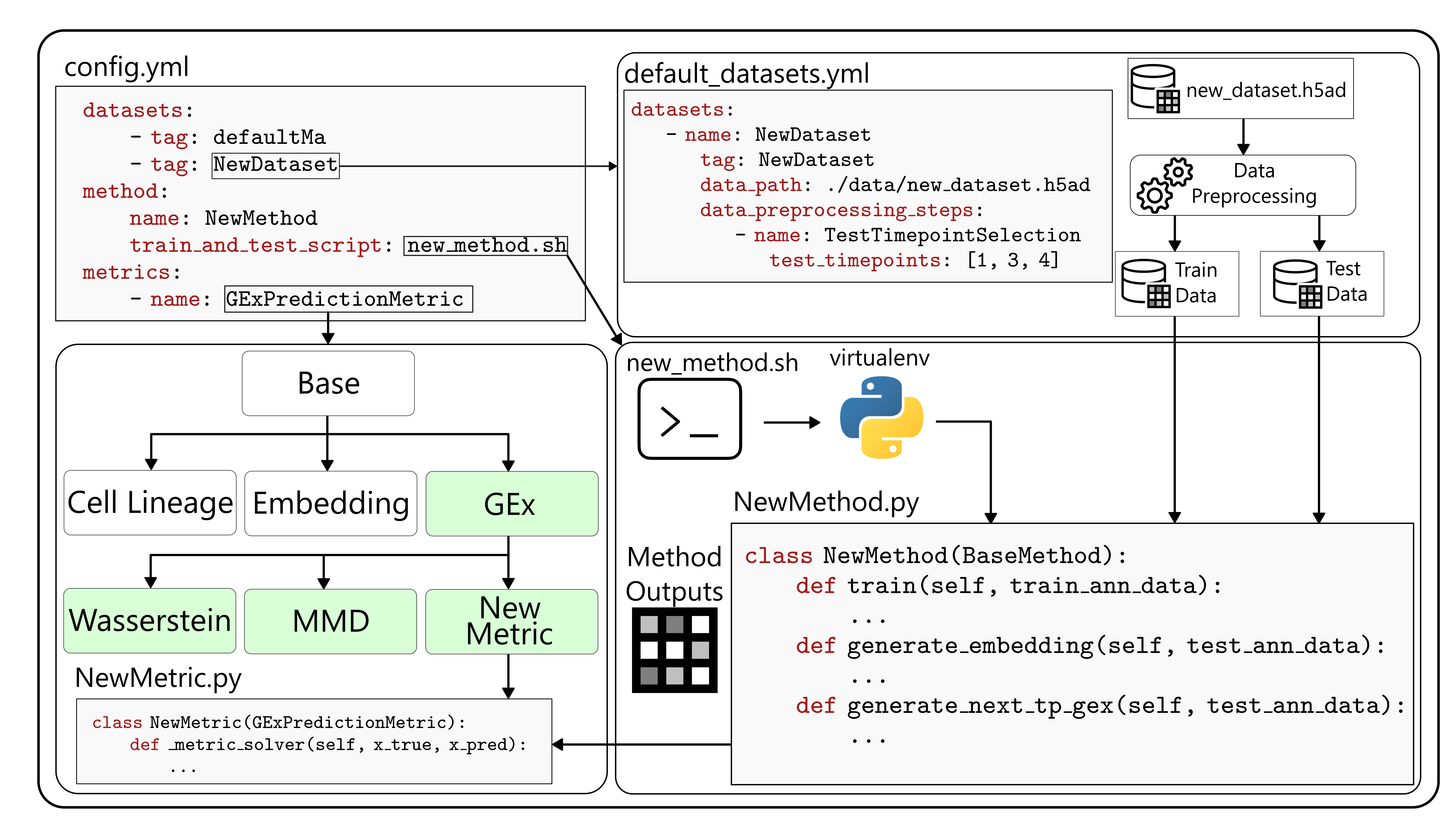
To achieve a smooth user experience, I made sure that users only need to include what’s minimally needed from them, which is:
- A configuration file that determines what the experiment will look like.
- Classes for each method that only require the necessary information for overriding its training and model output generation.
- Dataset classes and preprocessing steps for those datasets.
- Metric classes, which are ordered in a hierarchical tree.
For more information, read the preprint here: https://www.biorxiv.org/content/10.64898/2026.03.16.712069v1 and explore the GitHub: https://github.com/li-lab-mcgill/scTimeBench.